The Ethereal Landscape of the Interphase Nucleus
The nucleus of a eukaryotic cell, a bustling metropolis of genetic information, undergoes a profound transformation during the interphase stage of the cell cycle. While mitosis is characterized by the dramatic condensation of chromosomes into visible, rod-like structures, interphase presents a far more diffused and intricate picture. Within this quiescent period, the vast expanse of the cell’s DNA, meticulously organized and packaged, exists not as discrete chromosomes, but as a highly dynamic and complex network. This uncoiled, stringy DNA, a testament to the cell’s readiness for both replication and transcription, is collectively known as chromatin.
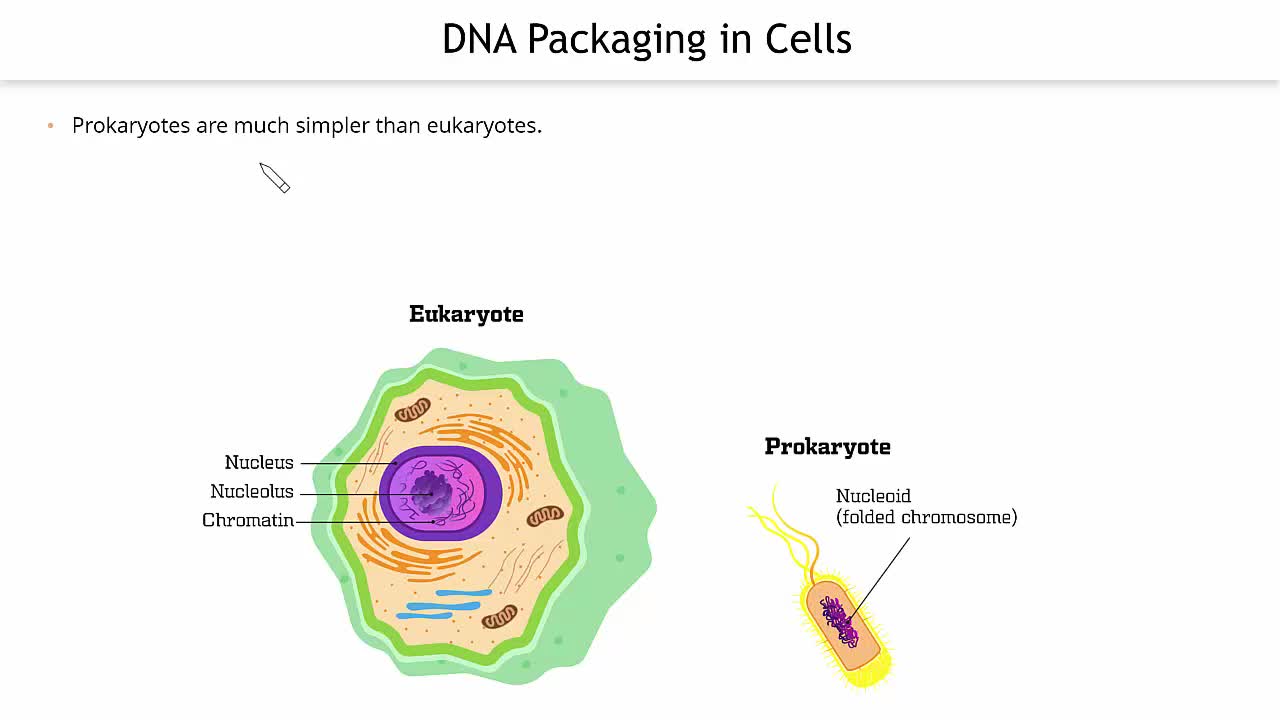
Chromatin is not simply a haphazard jumble of genetic material. Instead, it represents a sophisticated architectural marvel, designed to manage the immense length of DNA efficiently. Imagine the entire genome of a human cell, if stretched end-to-end, would measure approximately two meters in length. To fit this colossal molecule within the microscopic confines of the nucleus – a space typically only a few micrometers in diameter – requires an extraordinary level of compaction. This compaction is achieved through a hierarchical organization, where DNA is progressively wound and folded around specialized proteins.
During interphase, this organization is not static. The chromatin fiber undergoes continuous remodeling, allowing specific regions of the DNA to be accessed for essential cellular processes. This dynamic nature is crucial, as it dictates gene expression, DNA replication, and repair. The appearance of chromatin in interphase is often described as a “beads-on-a-string” model, where the “beads” are nucleosomes – the fundamental units of DNA packaging – and the “string” is the DNA molecule itself. However, this analogy, while helpful, simplifies a far more intricate reality. The chromatin fiber further folds and loops, creating a three-dimensional structure within the nucleus that is both organized and accessible.
Chromatin: The Building Blocks of Interphase DNA
The term “chromatin” itself is derived from the Greek words “chroma” (color) and “soma” (body), a nod to its characteristic staining properties when viewed under a microscope. While during mitosis, condensed chromosomes stain intensely, the more dispersed chromatin of interphase appears as a less intensely colored, granular material within the nucleus. This difference in staining reflects the level of DNA condensation.
At its most basic level, chromatin is composed of DNA and proteins. The primary proteins involved in DNA packaging are called histones. These are small, positively charged proteins that form octamers – structures consisting of eight histone subunits. The negatively charged DNA molecule wraps around these histone octamers, forming nucleosomes. Each nucleosome consists of approximately 147 base pairs of DNA wrapped around a histone core. This fundamental unit is the crucial first step in compacting the DNA.
The DNA wrapped around a histone octamer forms the “beads” in the “beads-on-a-string” model. The DNA linker between these nucleosomes is also bound by another histone, H1, which helps to further condense the chromatin fiber. This coiling of DNA around histones reduces the length of the DNA molecule by approximately sevenfold. However, this is just the beginning of the hierarchical packaging process.
Beyond the nucleosome level, the “beads-on-a-string” structure undergoes further folding. The 10-nanometer chromatin fiber, composed of nucleosomes, coils and folds to form a thicker, 30-nanometer fiber, often referred to as the “solenoid” model. This 30-nanometer fiber is thought to be formed by the coiling of the nucleosome fiber, with the help of histone H1. This level of organization further compacts the DNA, reducing its length by another factor.
The 30-nanometer fiber then loops and folds onto itself, forming larger structures. These loops are anchored to a protein scaffold, creating a more complex three-dimensional organization within the nucleus. This looping and anchoring process is crucial for regulating gene expression, as it allows for the specific folding and unfolding of DNA regions, making them accessible or inaccessible to the cellular machinery involved in transcription.
While the term “chromatin” encompasses all the DNA and proteins within the nucleus during interphase, it’s important to distinguish between different states of chromatin condensation. These states play a critical role in regulating gene activity.
Euchromatin: The Active Gene Territory
Within the diffuse mass of interphase chromatin, two main types can be distinguished based on their density and transcriptional activity: euchromatin and heterochromatin. Euchromatin is the less condensed, more transcriptionally active form of chromatin. It appears as a lighter, more dispersed material under the microscope.
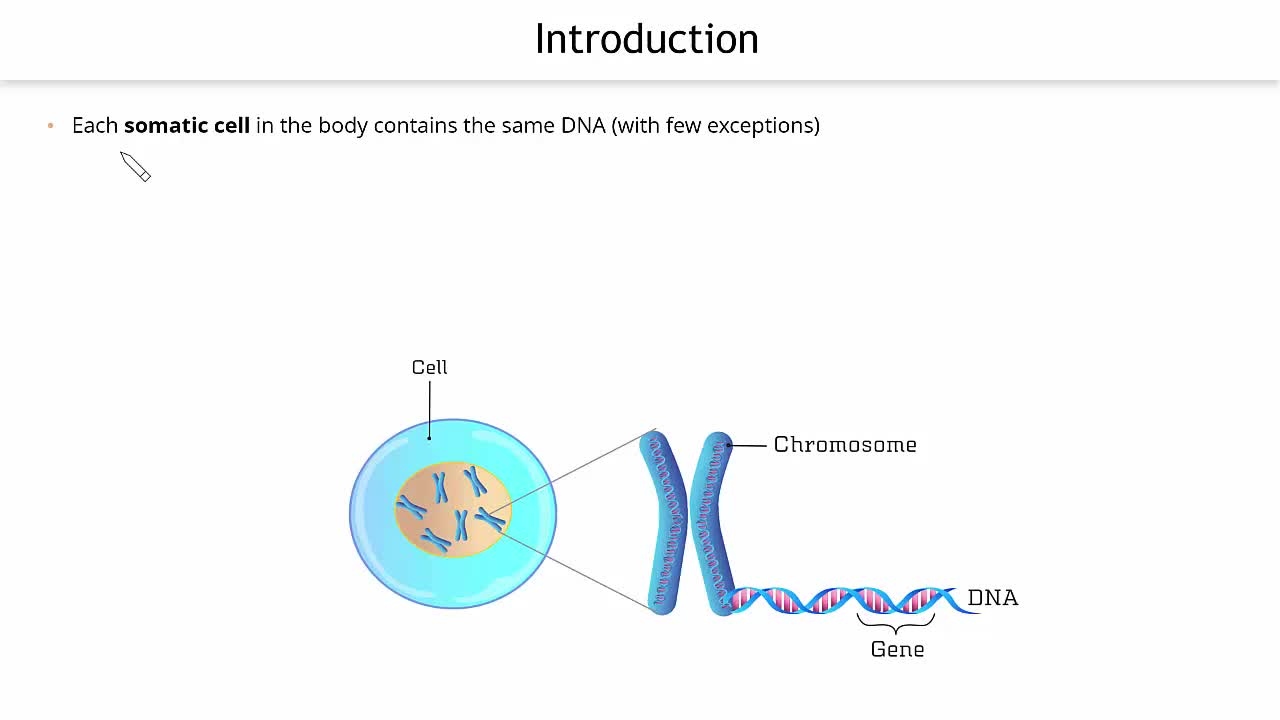
Euchromatin is where the majority of the cell’s genes reside and are actively being expressed. The DNA in euchromatin is more accessible to the molecular machinery responsible for transcription, such as RNA polymerase and transcription factors. This accessibility is facilitated by the looser packing of the DNA, allowing these proteins to bind to specific DNA sequences and initiate gene expression.
The dynamic nature of euchromatin is a hallmark of cellular activity. During interphase, specific regions of euchromatin can be further decondensed or condensed in response to cellular signals, allowing for fine-tuning of gene expression. This dynamic remodeling is often mediated by epigenetic modifications, such as the acetylation of histone tails. Histone acetylation generally leads to a loosening of chromatin structure, promoting gene expression.
Heterochromatin: The Silent Archives
In contrast to euchromatin, heterochromatin is the more condensed, transcriptionally inactive form of chromatin. It stains more darkly and appears as denser, more compact regions within the nucleus. Heterochromatin can be further subdivided into two types: constitutive heterochromatin and facultative heterochromatin.
Constitutive heterochromatin is permanently condensed and is typically found in specific regions of chromosomes, such as centromeres and telomeres. These regions often contain repetitive DNA sequences and play important roles in chromosome structure and stability. Because of its condensed nature, the genes within constitutive heterochromatin are generally not expressed.
Facultative heterochromatin, on the other hand, can switch between a condensed and decondensed state. It represents regions of the genome that are transcriptionally silenced at certain times or in certain cell types but can become active under specific conditions. For example, in female mammals, one of the two X chromosomes is largely inactivated and exists as facultative heterochromatin, known as a Barr body. This process of X-inactivation ensures that females, like males, have a balanced dosage of X-linked genes.
The presence of both euchromatin and heterochromatin within the interphase nucleus highlights the sophisticated regulatory mechanisms that govern gene expression and maintain genome integrity. This dynamic interplay between open and closed chromatin states allows the cell to control which genes are turned on or off at any given time, ensuring proper cellular function and development.

The Dynamic Nature of Interphase DNA Packaging
The “stringy DNA” of interphase chromatin is not a static entity. Its structure and organization are subject to constant regulation, a process crucial for all nuclear functions. This dynamic nature is mediated by a complex interplay of proteins and post-translational modifications.
One of the primary mechanisms driving chromatin dynamics is the activity of chromatin remodeling complexes. These are multi-protein machines that can alter the structure of nucleosomes, including sliding them along the DNA, ejecting them, or restructuring them. By repositioning nucleosomes, chromatin remodelers can expose or hide specific DNA sequences, thereby controlling access for transcription factors and other regulatory proteins.
Epigenetic modifications play a pivotal role in defining the different states of chromatin. These modifications are chemical alterations to DNA or histone proteins that do not change the underlying DNA sequence but can significantly influence gene expression. Histone modifications, such as acetylation, methylation, phosphorylation, and ubiquitination, are particularly important.
Histone acetylation, as mentioned earlier, generally loosens chromatin structure, making DNA more accessible and promoting gene transcription. Conversely, histone deacetylation leads to chromatin condensation and gene silencing. Histone methylation can have varied effects depending on the specific amino acid and the number of methyl groups added. For example, methylation of certain lysine residues on histone H3 can be associated with gene activation, while methylation of others can lead to gene repression.
These epigenetic marks are not random; they are dynamically added and removed by specific enzymes, creating a “code” that directs the assembly of various protein complexes involved in gene regulation. This “histone code” hypothesis suggests that combinations of histone modifications can dictate the functional state of a particular region of chromatin.
Furthermore, the three-dimensional organization of chromatin within the nucleus is also highly dynamic and plays a significant role in gene regulation. Regions of the genome that are actively transcribed or replicated are often found in specific sub-nuclear compartments. For instance, actively transcribed genes are frequently located in the interior of the nucleus, while heterochromatic regions are often found at the nuclear periphery. The formation of chromatin loops, anchored by proteins such as CTCF, can bring distant regulatory elements into close proximity with their target genes, facilitating or inhibiting transcription.
This intricate and dynamic organization of chromatin during interphase ensures that the cell can efficiently manage its vast genetic information, respond to environmental cues, and carry out essential processes like DNA replication and repair, all while maintaining the integrity of the genome. The uncoiled stringy DNA, in its chromatin form, is the very essence of a living cell’s ongoing narrative of life and function.
