Immunoprecipitation (IP) is a powerful laboratory technique used to isolate and purify a specific protein from a complex mixture of cellular proteins. It leverages the exquisite specificity of antibody-antigen interactions to selectively pull down, or precipitate, the target protein of interest. This allows researchers to study its properties, interactions, and abundance, providing crucial insights into cellular processes, disease mechanisms, and drug targets. While the underlying principle is straightforward, the execution involves several critical steps and considerations, making it a cornerstone of modern molecular biology research.
The Principle of Antibody-Antigen Recognition
At the heart of immunoprecipitation lies the highly specific binding between an antibody and its corresponding antigen. Antibodies are Y-shaped proteins produced by the immune system that are designed to recognize and bind to specific molecular structures, known as epitopes, on foreign substances or abnormal cells. In the context of IP, the antigen is typically the protein that a researcher wishes to study.
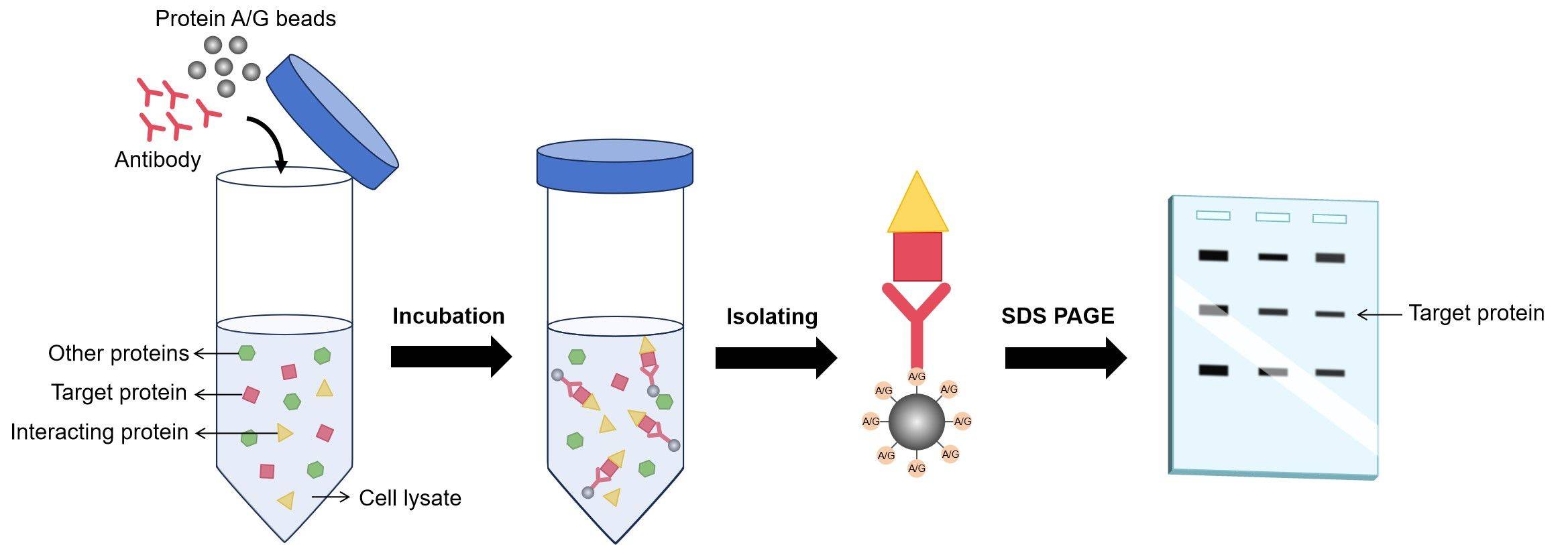
When an antibody is mixed with a sample containing the target protein, the antibody’s antigen-binding sites will selectively attach to the epitopes on the target protein. This binding is remarkably precise, meaning that a well-designed antibody will only bind to its intended target and not to other, unrelated proteins that may be present in the sample. This specificity is what makes IP such a valuable tool for isolating individual proteins from the noisy background of a cellular lysate.
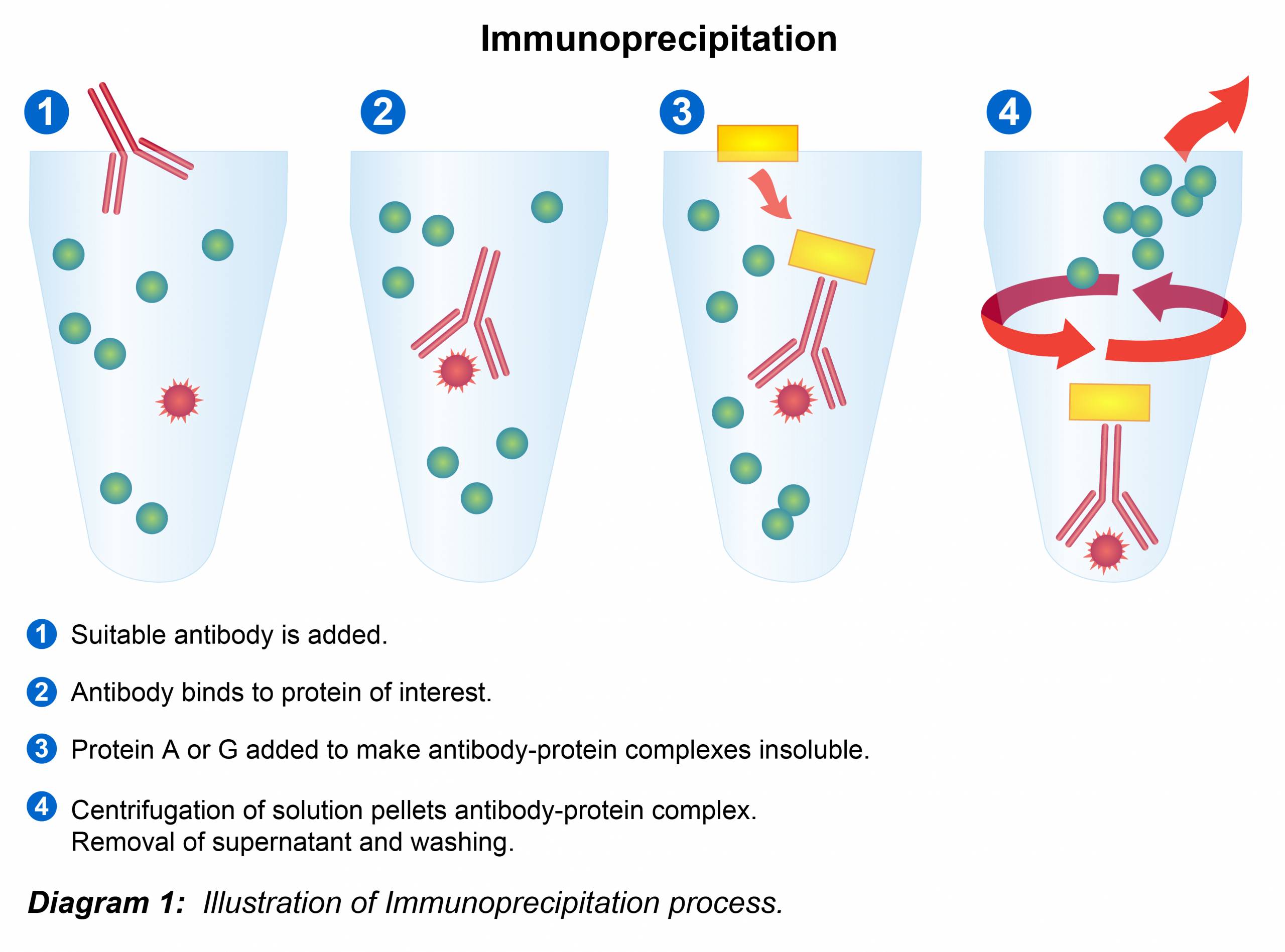
The process begins with the preparation of a cell or tissue lysate, which is a complex soup of all the proteins and other molecules within the cells. This lysate is then incubated with a specific antibody that has been chosen to bind to the protein of interest. Often, this antibody is conjugated, or attached, to a solid support, such as magnetic beads or agarose beads. These beads provide a convenient way to physically separate the antibody-protein complex from the rest of the lysate.
After the incubation period, which allows ample time for antibody-antigen binding, the beads (along with the bound target protein and antibody) are separated from the supernatant containing the unbound cellular components. This separation can be achieved through various methods, such as centrifugation or magnetic separation, depending on the type of beads used. The beads are then washed rigorously to remove any non-specifically bound molecules, ensuring that the isolated protein is as pure as possible. Finally, the bound target protein is eluted from the beads, typically by changing the pH or using a chaotropic agent, releasing it into a soluble form for further analysis.
Key Components and Reagents in Immunoprecipitation
Successful immunoprecipitation relies on the careful selection and use of several key components and reagents. Each plays a critical role in ensuring the efficiency and specificity of the isolation process.
Antibodies: The Specific Probes
The choice of antibody is paramount in IP. Antibodies can be monoclonal (produced from a single B cell clone, recognizing a single epitope) or polyclonal (produced from multiple B cell clones, recognizing multiple epitopes on the same antigen). Monoclonal antibodies generally offer higher specificity, reducing the likelihood of cross-reactivity with other proteins. Polyclonal antibodies, on the other hand, can sometimes provide stronger signal due to binding to multiple epitopes, which can be advantageous if the target protein is present at low concentrations or if its native conformation is not optimally recognized by a single epitope.
The antibody must be validated for its ability to bind to the target protein in its native or denatured state, depending on the experimental conditions. It’s crucial that the antibody recognizes an epitope that is accessible in the cellular environment and that is not masked by protein-protein interactions.
Solid Supports: Beads for Capture
To facilitate the separation of the antibody-antigen complex from the solution, the antibody is typically immobilized on a solid support, most commonly in the form of beads.
- Agarose Beads: These are widely used and offer a good balance of binding capacity and ease of handling. They are often cross-linked to improve their stability.
- Magnetic Beads: These are becoming increasingly popular due to the ease and speed of separation. The magnetic properties allow for rapid collection of the beads and bound complexes using a magnet, eliminating the need for centrifugation and reducing potential sample loss. They can be coated with various binding agents, such as Protein A or Protein G, which bind to the Fc region of antibodies.
- Sepharose Beads: Similar to agarose, these are also commonly used and offer good binding capacity.
The choice of bead material and its coating (e.g., Protein A, Protein G, or directly conjugated antibodies) depends on the type of antibody being used and the specific experimental requirements. Protein A and Protein G are bacterial proteins that have a high affinity for the Fc region of antibodies, allowing them to capture a wide range of antibodies efficiently.
Lysis Buffers: Preserving Protein Integrity
The lysis buffer is designed to break open cells and release their contents while simultaneously preserving the integrity of the target protein. This involves solubilizing cellular membranes and denaturing intracellular structures. Lysis buffers typically contain:
- Detergents: Such as Triton X-100, NP-40, or SDS, to disrupt cell membranes and solubilize proteins. The type and concentration of detergent are critical as harsh detergents can denature proteins and disrupt protein-protein interactions.
- Salts: Like NaCl, to control ionic strength and aid in protein solubilization.
- Buffering agents: Such as Tris-HCl, to maintain a stable pH, which is essential for protein stability and antibody-antigen binding.
- Protease Inhibitors: A cocktail of inhibitors is usually added to prevent the degradation of proteins by endogenous proteases released during cell lysis.
- Phosphatase Inhibitors: If the study involves post-translational modifications like phosphorylation, phosphatase inhibitors are included.
The composition of the lysis buffer can significantly impact the success of IP, especially when studying transient protein interactions or proteins that are sensitive to harsh conditions.
Washing Buffers: Eliminating Non-Specific Binding
After capturing the antibody-protein complex, thorough washing is essential to remove any proteins or molecules that have bound non-specifically to the beads or antibody. Washing buffers are typically similar to lysis buffers but often with a slightly lower detergent concentration to avoid dissociating the specifically bound target protein. Multiple washes are usually performed to ensure the highest possible purity of the immunoprecipitated sample.
Elution Buffers: Releasing the Target Protein
Once the target protein is bound to the antibody on the beads, it needs to be released for subsequent analysis. Elution buffers are designed to disrupt the antibody-antigen interaction. Common elution strategies include:
- Low pH Elution: Using buffers with a pH around 2-3 can disrupt antibody-antigen binding. The eluted sample is then immediately neutralized to prevent protein denaturation.
- High pH Elution: Less common but can also be effective.
- Chaotropic Agents: Solutions containing agents like sodium dodecyl sulfate (SDS) or urea can denature proteins and disrupt the binding.
- Competitive Elution: Adding excess free antigen or a peptide corresponding to the epitope recognized by the antibody can outcompete the binding of the target protein.
The choice of elution method depends on the stability of the target protein and the downstream application.
Types of Immunoprecipitation Techniques
While the fundamental principle of IP remains the same, several variations of the technique have been developed to suit different research questions and to enhance specific aspects of the process.
Standard Immunoprecipitation (IP)
This is the most basic form of IP, where a specific antibody is used to precipitate a target protein from a cell lysate. The precipitated protein can then be analyzed by Western blotting, mass spectrometry, or other methods to confirm its identity, assess its abundance, or investigate its modifications.
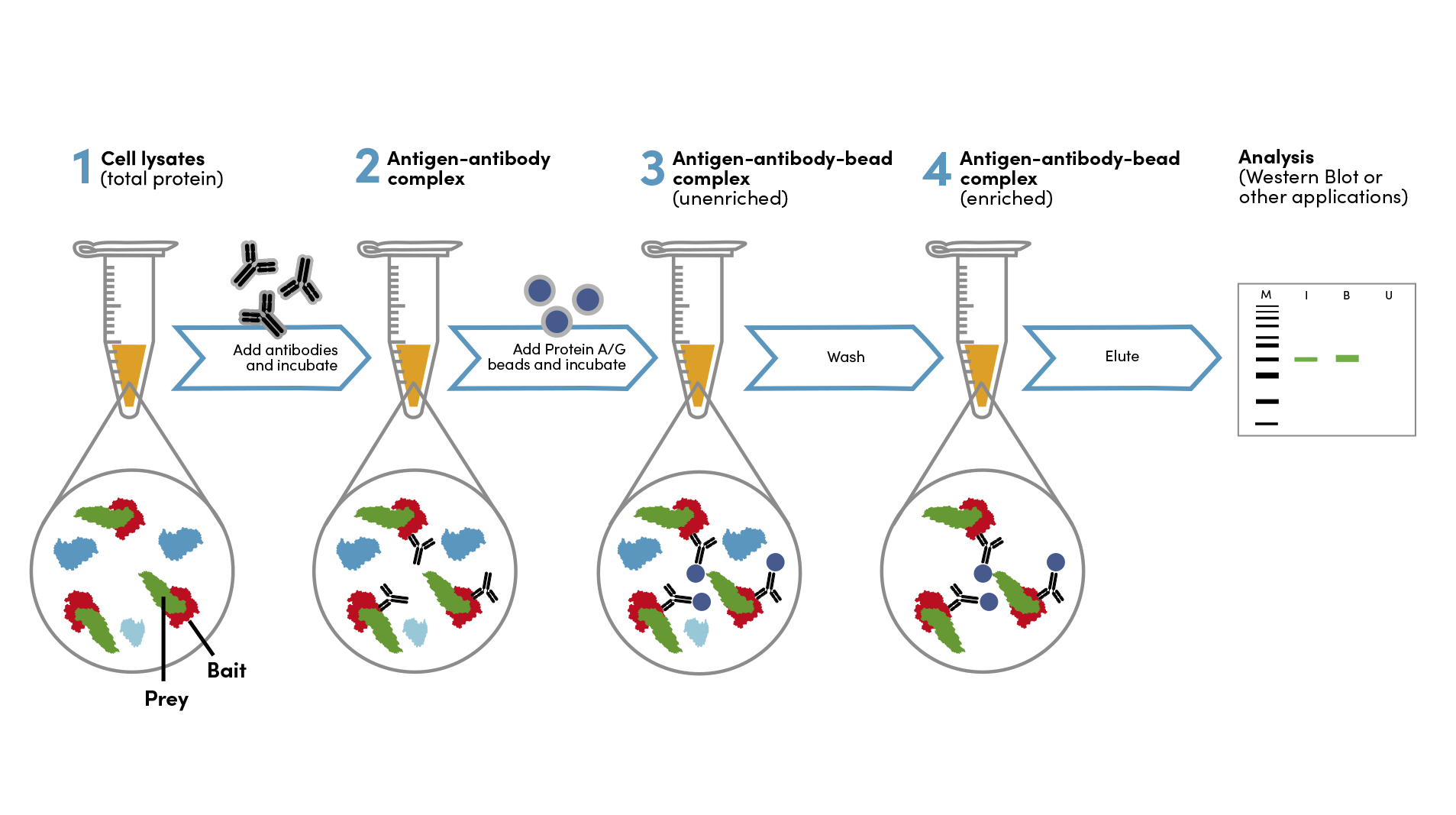
Co-Immunoprecipitation (Co-IP)
Co-IP is a widely used technique to identify protein-protein interactions. In this method, a cell lysate is incubated with an antibody against one of the proteins suspected to be interacting (the bait protein). If a physical interaction exists, the interacting partner (the prey protein) will be pulled down along with the bait protein. The precipitated complex is then analyzed, typically by Western blotting using an antibody against the prey protein. This allows researchers to infer that the two proteins interact directly or indirectly within the cellular context.
Chromatin Immunoprecipitation (ChIP)
ChIP is specifically used to study the interaction of proteins, such as transcription factors or histone modifications, with DNA. In ChIP, the cellular material is first cross-linked to fix protein-DNA complexes. The DNA is then fragmented, and an antibody against the protein of interest is used to precipitate the protein-DNA complexes. After reversing the cross-linking and purifying the DNA, it can be analyzed by techniques like PCR or sequencing to identify the DNA regions bound by the protein.
Immunoprecipitation of RNA (RIP)
RNA immunoprecipitation (RIP) is analogous to ChIP but focuses on isolating RNA-binding proteins and their associated RNA molecules. After IP of the protein of interest, the co-precipitated RNA is isolated and can be analyzed to identify the specific RNAs that are bound by the protein. This is crucial for understanding RNA regulation, processing, and localization.
Immunoprecipitation of Post-Translational Modifications (PTM-IP)
This technique involves using antibodies that specifically recognize post-translational modifications on proteins, such as phosphorylation, ubiquitination, or acetylation. By precipitating proteins with these modifications, researchers can study the dynamics and functional consequences of these modifications within the cell. For example, phospho-IP can be used to identify kinases and their substrates or to understand signaling pathways.
Applications and Significance of Immunoprecipitation
The versatility of immunoprecipitation has led to its widespread application across numerous fields of biological research.
Elucidating Protein Function and Pathways
By isolating and characterizing specific proteins, researchers can gain a deeper understanding of their roles in cellular processes. IP is instrumental in identifying components of signaling pathways, metabolic enzymes, structural proteins, and regulatory molecules. Studying the abundance and modifications of these proteins under different conditions can reveal how they contribute to normal cellular function and how their dysregulation leads to disease.
Investigating Protein-Protein Interactions
As highlighted with Co-IP, understanding how proteins interact with each other is fundamental to deciphering the molecular architecture and functional organization of cells. IP allows for the mapping of interaction networks, revealing the dynamic assembly of protein complexes that carry out specific cellular tasks. This is vital for drug discovery, as many therapeutic targets are proteins involved in critical interaction hubs.
Studying Disease Mechanisms
Many diseases, including cancer, neurodegenerative disorders, and infectious diseases, are characterized by aberrant protein function or interactions. IP can be used to identify disease-specific protein alterations, such as over- or under-expression, abnormal modifications, or novel interaction partners. This knowledge is essential for developing diagnostic markers and targeted therapies.
Drug Discovery and Development
Identifying how drugs affect protein targets and their interactions is a critical aspect of drug discovery. IP can be used to:
- Validate drug targets: Confirming that a drug candidate indeed binds to its intended target and alters its activity.
- Characterize off-target effects: Identifying unintended interactions of a drug with other proteins, which can lead to side effects.
- Monitor drug efficacy: Assessing changes in protein levels or interactions in response to drug treatment.
In conclusion, immunoprecipitation is a sophisticated and indispensable tool in the molecular biologist’s arsenal. Its ability to selectively isolate and analyze specific proteins from complex biological samples underpins a vast array of research endeavors, driving advancements in our understanding of life at the molecular level and paving the way for novel therapeutic interventions.
