Restriction enzymes, often referred to as molecular scissors, are fundamental tools in molecular biology, playing a crucial role in recombinant DNA technology and genetic engineering. Their primary function is to recognize specific nucleotide sequences within a DNA molecule and cleave the DNA at precise locations. This ability to cut DNA at predictable sites is the cornerstone of many advanced biotechnological applications.
The Mechanism of Restriction Enzymes
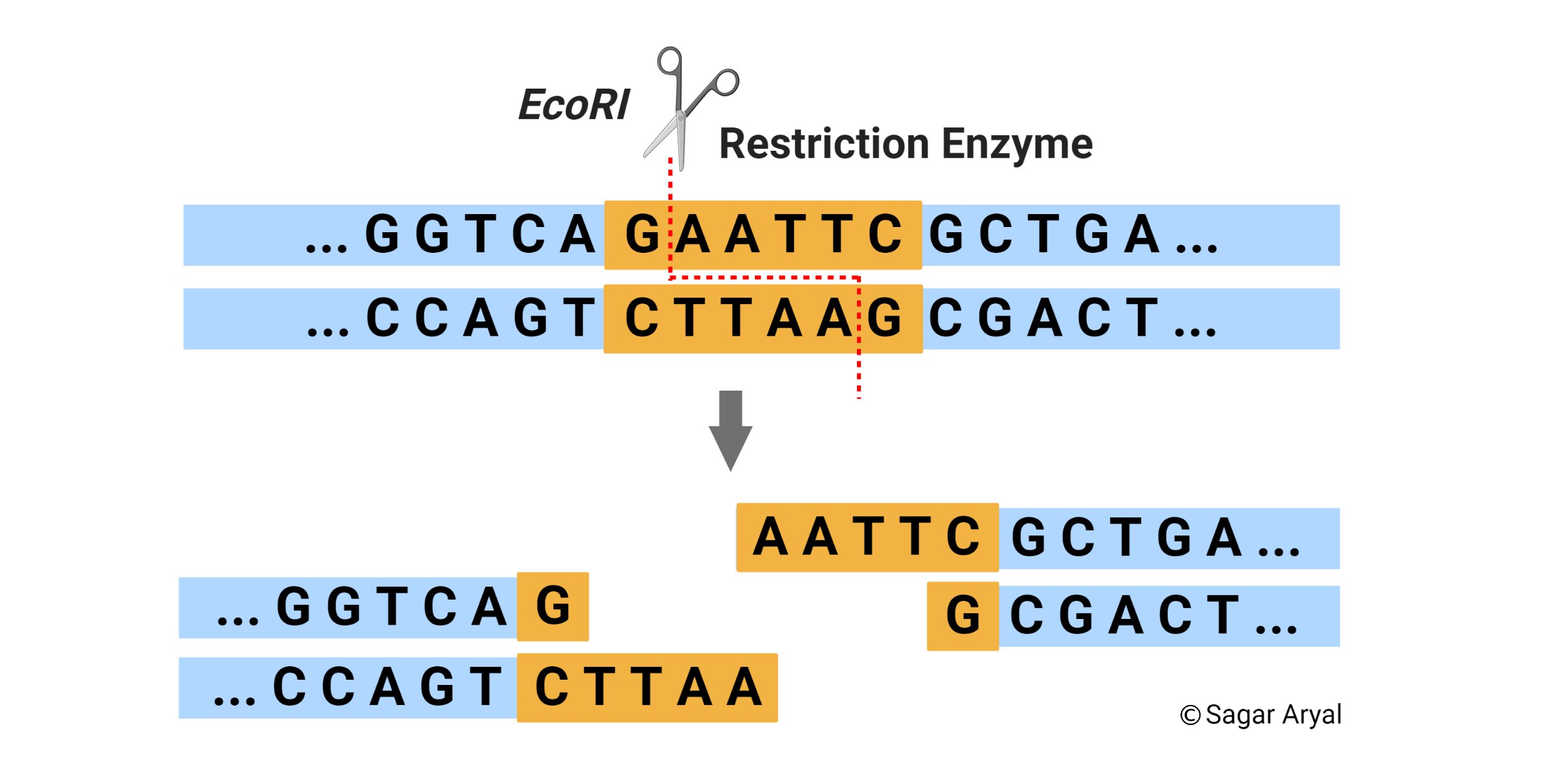
The specificity of a restriction enzyme stems from its recognition sequence, also known as a restriction site. These sites are typically short, palindromic sequences, meaning they read the same forwards and backwards on opposite strands of the DNA. For example, the restriction enzyme EcoRI recognizes the sequence GAATTC. When EcoRI encounters this sequence, it binds to the DNA and catalyzes the hydrolysis of the phosphodiester bond between guanine (G) and adenine (A) on one strand, and between adenine (A) and guanine (G) on the complementary strand.
Types of Restriction Enzymes
Restriction enzymes are classified into three main types based on their structure, the nature of their cleavage, and their cofactor requirements.
Type I Restriction Enzymes
Type I restriction enzymes possess both restriction and modification activities. They recognize specific DNA sequences and can cleave DNA at sites that are relatively distant from the recognition sequence, often hundreds or thousands of base pairs away. These enzymes require ATP and S-adenosylmethionine (SAM) as cofactors for their activity. Due to the variability in their cleavage sites, Type I enzymes are less commonly used in routine molecular cloning applications.
Type II Restriction Enzymes
Type II restriction enzymes are the most widely used and extensively studied type. They recognize specific DNA sequences and cleave the DNA directly within or adjacent to these recognition sites. A key characteristic of Type II enzymes is that their recognition sites are generally palindromic and are typically 4 to 8 base pairs in length. They do not require ATP or SAM for their activity, relying solely on the presence of magnesium ions (Mg2+). This predictable and precise cleavage makes them invaluable for applications like gene cloning, DNA mapping, and Southern blotting.
Type II restriction enzymes can be further subdivided into several subclasses (Type IIA, IIB, IIC, IID, IIE, IIF, IIH, IIK, etc.) based on their subunit composition, cofactor requirements, and cleavage patterns. However, for practical molecular biology applications, understanding the basic Type II mechanism is usually sufficient.
Type III Restriction Enzymes
Type III restriction enzymes also possess both restriction and modification activities. They recognize specific DNA sequences, typically 5 to 7 base pairs long, and cleave the DNA at a site usually located about 20-25 base pairs downstream of the recognition sequence. These enzymes require ATP and Mg2+ for their activity and are sensitive to SAM. While less common than Type II enzymes, Type III enzymes are used in specific applications, particularly in studies of DNA repair and recombination.
The Role of Restriction Enzymes in Molecular Biology
The ability of restriction enzymes to cut DNA at specific sites has revolutionized molecular biology. They are essential tools for manipulating DNA and constructing recombinant molecules.
Gene Cloning
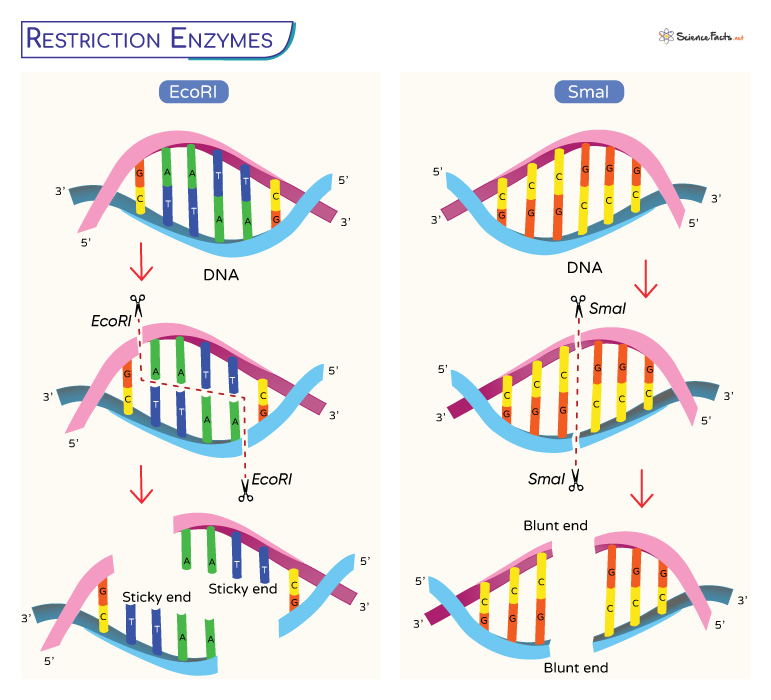
Perhaps the most prominent application of restriction enzymes is in gene cloning. To insert a gene of interest into a plasmid vector, both the gene and the plasmid are typically cut with the same restriction enzyme. This generates compatible “sticky ends” or “blunt ends” on both the DNA fragment containing the gene and the linearized plasmid. Sticky ends are short, single-stranded overhangs that can base-pair with complementary overhangs, facilitating the joining of the two DNA molecules by DNA ligase. Blunt ends lack overhangs and are ligated together through a blunt-end ligation process. Once the gene is inserted into the plasmid, the recombinant plasmid can be introduced into host cells, such as bacteria, where it can be replicated and expressed.
DNA Mapping
Restriction enzymes are also used to create restriction maps of DNA molecules. By digesting a DNA molecule with one or more restriction enzymes and analyzing the sizes of the resulting fragments using gel electrophoresis, scientists can determine the relative positions of restriction sites within the DNA. This information is crucial for understanding the organization of genes, identifying mutations, and sequencing DNA.
Southern Blotting
Southern blotting is a technique used to detect specific DNA sequences within a DNA sample. In this process, DNA is digested with restriction enzymes, separated by size using gel electrophoresis, and then transferred to a membrane. The membrane is then hybridized with a labeled probe that is complementary to the target DNA sequence. Restriction enzyme digestion is a critical first step in this technique, as it breaks down the large genomic DNA into manageable fragments that can be analyzed.
Genetic Engineering and Biotechnology
Beyond cloning and mapping, restriction enzymes are integral to a vast array of genetic engineering applications. They enable the precise excision and insertion of DNA segments, which is fundamental for:
- Creating Genetically Modified Organisms (GMOs): Introducing desirable traits into plants, animals, or microorganisms.
- Gene Therapy: Delivering therapeutic genes into cells to treat genetic disorders.
- Developing Diagnostic Tools: Creating DNA-based tests for diseases and genetic predispositions.
- Producing Recombinant Proteins: Manufacturing therapeutic proteins like insulin and growth hormone in large quantities.
- Synthetic Biology: Constructing novel biological systems with engineered functions.
Practical Considerations for Using Restriction Enzymes
Working with restriction enzymes requires careful attention to several factors to ensure optimal activity and successful experimental outcomes.
Buffer Conditions
Each restriction enzyme has an optimal buffer in which it exhibits maximum activity. These buffers typically contain salts (like NaCl and KCl), magnesium ions (Mg2+), and a buffering agent (such as Tris-HCl) to maintain a stable pH. It is crucial to use the correct buffer recommended by the enzyme manufacturer.
Temperature
Most restriction enzymes function optimally at 37°C. However, some enzymes may have different optimal temperatures. Incubation times are usually between 15 minutes and several hours, depending on the enzyme and the amount of substrate DNA.
Enzyme Purity and Activity
The purity and activity of the restriction enzyme are paramount. High-quality enzymes from reputable suppliers are essential to avoid non-specific digestion or the presence of contaminating nucleases that could degrade the DNA. Unit definitions, which represent the amount of enzyme required to digest a specific amount of DNA in a given time, are standardized to ensure consistent results.
Star Activity
Under certain conditions, restriction enzymes can exhibit “star activity,” meaning they cleave DNA at sequences that are not their intended recognition sites. Star activity can be induced by factors such as:
- High enzyme concentration: Using an excessive amount of enzyme relative to the substrate DNA.
- Low salt concentration: Performing the digestion in a buffer with insufficient salt.
- High pH: Working at a pH significantly higher than the enzyme’s optimum.
- Presence of organic solvents: Certain organic solvents can promote star activity.
- Extended incubation times: Prolonged incubation can sometimes lead to non-specific cleavage.
To avoid star activity, it is essential to adhere to the manufacturer’s recommended reaction conditions.

Isoschizomers and Neoschizomers
Isoschizomers are different restriction enzymes that recognize and cleave the same DNA sequence. Neoschizomers are enzymes that recognize similar, but not identical, sequences and may produce different cleavage patterns. The availability of multiple enzymes that cut at the same site provides flexibility in experimental design and can be useful if one enzyme is unavailable or has undesirable properties.
In conclusion, restriction enzymes are indispensable tools in molecular biology. Their ability to precisely cleave DNA at defined recognition sequences underpins a vast array of genetic manipulation techniques, from fundamental gene cloning to advanced applications in genetic engineering and synthetic biology. Understanding their mechanism, types, and optimal usage is crucial for any researcher working with DNA.
