The intricate process of DNA replication is fundamental to life itself, ensuring the faithful transmission of genetic information from one generation to the next. At the heart of this molecular machinery lies a family of enzymes known as polymerases, specifically DNA polymerases. These remarkable biological catalysts are the architects and builders of new DNA strands, meticulously assembling nucleotides into precise sequences. Without their coordinated action, the duplication of our genetic blueprint would be impossible, leading to cellular dysfunction and, ultimately, the cessation of life. This exploration delves into the multifaceted role of polymerase in DNA replication, dissecting its mechanisms, identifying key players, and appreciating its indispensable contribution to biological continuity.
The Core Function: Synthesizing New DNA Strands
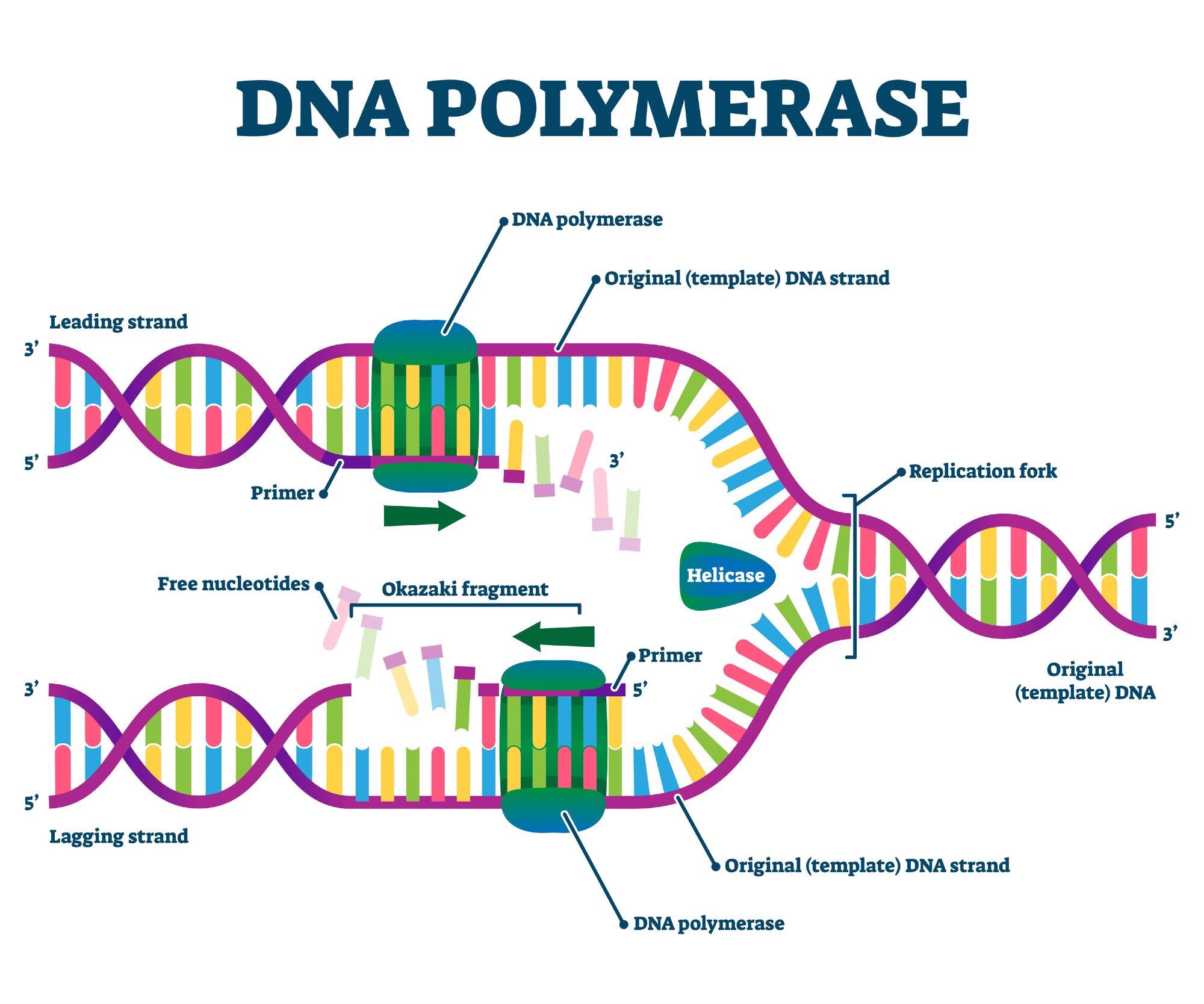
The primary role of DNA polymerase is to catalyze the formation of phosphodiester bonds, linking deoxynucleoside triphosphates (dNTPs) together to form a new DNA strand. This process is remarkably specific, guided by the template strand of the parental DNA molecule. DNA polymerase does not initiate DNA synthesis de novo; rather, it requires a pre-existing strand of nucleotides, typically an RNA primer synthesized by an enzyme called primase, to begin its work. This primer provides a free 3′-hydroxyl (OH) group to which the polymerase can add the first deoxyribonucleotide.
The polymerase then reads the template strand in the 3′ to 5′ direction and synthesizes the new complementary strand in the 5′ to 3′ direction. This directional synthesis is a crucial characteristic of DNA replication. For each nucleotide added to the growing chain, the polymerase cleaves two phosphate groups from the incoming dNTP, releasing energy that drives the polymerization reaction. The accuracy of this process is paramount, as even a single incorrect nucleotide incorporation can have profound consequences for the organism.
Mechanism of Nucleotide Addition
The active site of DNA polymerase is exquisitely designed to accommodate the incoming dNTP and the growing DNA strand. The incoming dNTP base-pairs with its complementary base on the template strand through hydrogen bonding (adenine with thymine, guanine with cytosine). This correct base pairing is essential for the polymerase to proceed. Once the correct dNTP is in place, the 3′-OH group of the last nucleotide in the growing chain attacks the alpha-phosphate of the incoming dNTP, forming a phosphodiester bond. This reaction results in the elongation of the new DNA strand by one nucleotide and the release of pyrophosphate (two phosphate groups).
Proofreading and Error Correction
While DNA polymerases are incredibly accurate, errors do occur. The spontaneous mutation rate in DNA replication is low but not zero. To combat this, many DNA polymerases possess an intrinsic proofreading activity. This 3′ to 5′ exonuclease activity allows the polymerase to detect and remove incorrectly incorporated nucleotides. If the polymerase adds a nucleotide that does not properly base-pair with the template, it can pause, reposition the mismatched nucleotide to its exonuclease site, and cleave it from the growing chain. This “back-editing” mechanism significantly reduces the error rate of DNA replication, ensuring the fidelity of genetic information transfer.
Key Players in the Replication Machinery
While the term “polymerase” often refers to the core catalytic enzyme, DNA replication is a complex process involving numerous proteins that work in concert with the polymerase. Understanding the roles of these accessory proteins is crucial to appreciating the overall efficiency and accuracy of DNA replication.
The Replication Fork
DNA replication proceeds bidirectionally from specific origins of replication. The replication fork is the Y-shaped structure where the parental DNA double helix is unwound and new strands are being synthesized. Several key enzymes operate at the replication fork to facilitate this process.
Helicase: Unwinding the Double Helix
Helicase enzymes are responsible for breaking the hydrogen bonds that hold the two strands of the DNA double helix together. This unwinding activity separates the parental DNA strands, making them accessible as templates for new DNA synthesis. Helicases move along the DNA, consuming ATP, and effectively “zipping” open the double helix.
Single-Strand Binding Proteins (SSBs): Stabilizing the Unwound Strands
Once the DNA strands are separated by helicase, they have a tendency to re-anneal or form secondary structures. Single-strand binding proteins (SSBs) bind to the exposed single strands of DNA, preventing them from re-forming a double helix and protecting them from degradation by nucleases. They also help to keep the DNA strands in a linear conformation, facilitating the action of other replication enzymes.

Primase: Initiating DNA Synthesis
As mentioned earlier, DNA polymerase cannot initiate synthesis on its own. Primase, an RNA polymerase, synthesizes short RNA primers (typically 5-10 nucleotides long) complementary to the template DNA strand. These RNA primers provide the essential 3′-OH group that DNA polymerase needs to begin adding deoxyribonucleotides.
DNA Ligase: Joining DNA Fragments
The synthesis of the lagging strand, which proceeds discontinuously, results in short fragments of DNA called Okazaki fragments. DNA ligase is an enzyme that seals the nicks between these fragments by forming a phosphodiester bond, thereby creating a continuous DNA strand.
Different Types of DNA Polymerases
In both prokaryotes and eukaryotes, there are multiple types of DNA polymerases, each with specialized roles in replication, repair, and other DNA-related processes.
Prokaryotic DNA Polymerases
Prokaryotes, such as E. coli, typically have three main DNA polymerases:
- DNA Polymerase I (Pol I): This enzyme plays a crucial role in removing RNA primers from Okazaki fragments and filling in the resulting gaps with DNA. It also has a significant 5′ to 3′ exonuclease activity that contributes to primer removal. Pol I is involved in DNA repair as well.
- DNA Polymerase II (Pol II): Primarily involved in DNA repair, particularly in response to DNA damage. It possesses 3′ to 5′ exonuclease activity for proofreading.
- DNA Polymerase III (Pol III): This is the primary replicative polymerase in E. coli. It is a highly processive enzyme, meaning it can synthesize long stretches of DNA without dissociating from the template. Pol III is responsible for the bulk of DNA synthesis during replication of both the leading and lagging strands. It also has 3′ to 5′ exonuclease activity for proofreading.
Eukaryotic DNA Polymerases
Eukaryotes have a more complex array of DNA polymerases, with at least 15 known types. The most important ones for nuclear DNA replication are:
- DNA Polymerase α (Pol α): This polymerase is part of a complex that also includes primase. It initiates DNA synthesis by synthesizing a short RNA primer followed by a short stretch of DNA nucleotides. Pol α has low processivity and lacks proofreading activity, so it is quickly replaced by other polymerases.
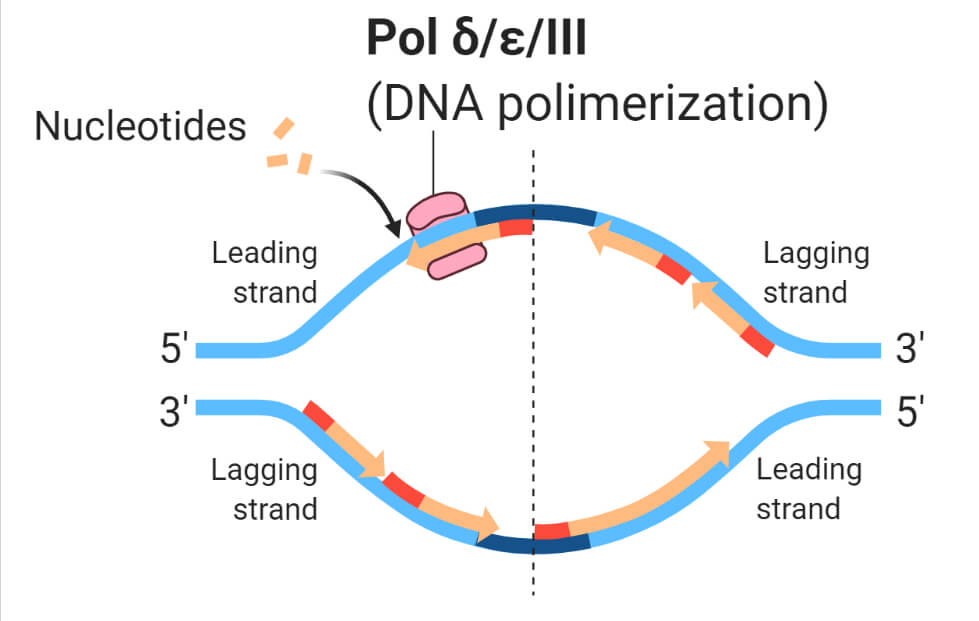
- DNA Polymerase δ (Pol δ): This is one of the main replicative polymerases in eukaryotes, responsible for synthesizing the lagging strand. It is highly processive and possesses 3′ to 5′ exonuclease proofreading activity.
- DNA Polymerase ε (Pol ε): The other major replicative polymerase, primarily responsible for synthesizing the leading strand. It is also highly processive and has robust proofreading activity. Pol ε is thought to be more involved in the fidelity of replication.
- DNA Polymerase β (Pol β): Primarily involved in DNA repair, specifically in base excision repair. It is a small, non-processive polymerase.
- Mitochondrial DNA Polymerase (Pol γ): This polymerase is responsible for replicating the DNA found within mitochondria.
The coordinated action of these various polymerases, along with other replication proteins, ensures that the entire genome is accurately duplicated before cell division.
The Significance of Polymerase Accuracy and Processivity
The fidelity of DNA replication is critical for maintaining the integrity of the genome. Errors in DNA replication can lead to mutations, which can have detrimental effects on cellular function and organismal health, potentially contributing to diseases like cancer and genetic disorders. The proofreading capabilities of DNA polymerases are a crucial safeguard against such errors.
Processivity, the ability of a polymerase to remain attached to the DNA template and continue synthesis for extended periods, is also vital for efficient replication. Highly processive polymerases can synthesize large segments of DNA without frequently dissociating, thus speeding up the overall replication process. Eukaryotic polymerases, particularly Pol δ and Pol ε, achieve high processivity through association with other proteins, such as the sliding clamp (PCNA), which acts like a ring that encircles the DNA and tethers the polymerase to it.
In conclusion, DNA polymerases are the indispensable workhorses of DNA replication. Their ability to read a template, select and incorporate complementary nucleotides, and proofread for errors forms the bedrock of accurate genetic inheritance. The diverse array of polymerases and their intricate coordination with other replication factors underscore the complexity and elegance of this fundamental biological process. Understanding the role of polymerase is not just an academic pursuit; it is a window into the very mechanisms that sustain life and ensure its continuation across generations.
