The intricate machinery of life relies on a constant supply of energy, and at the heart of this energy production lies adenosine triphosphate (ATP). This ubiquitous energy currency is synthesized by a remarkable molecular machine known as ATP synthase. But what precisely drives the incredibly efficient and complex process of ATP generation? The answer lies in a subtle yet powerful force: a concentration gradient. More specifically, it is an electrochemical gradient that fuels the rotary motor of ATP synthase, ultimately converting proton or sodium ion flow into the chemical energy of ATP.
ATP synthase is a marvel of biological engineering, found embedded within the membranes of mitochondria (in eukaryotes), chloroplasts (in plants and algae), and bacterial plasma membranes. Its structure is elegantly bipartite, consisting of two main components: the F₀ sector, embedded within the membrane, and the F₁ sector, protruding into the catalytic space. The F₀ sector acts as a proton or ion channel, while the F₁ sector houses the catalytic sites where ATP is actually synthesized. The magic happens when protons (or sometimes sodium ions) accumulate on one side of the membrane, creating a difference in concentration and charge – the electrochemical gradient. This gradient represents stored potential energy, analogous to water behind a dam.
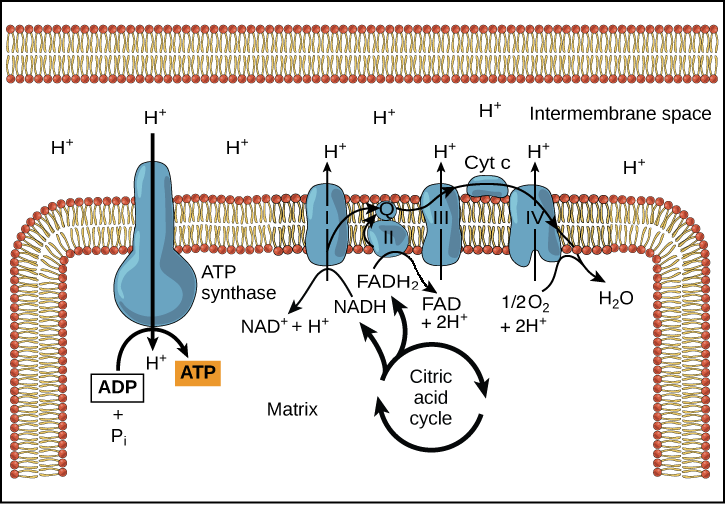
The Proton-Motive Force: Nature’s Battery
The primary driving force behind ATP synthesis, particularly in aerobic respiration and photosynthesis, is the proton-motive force (PMF). This concept encapsulates the combined effects of both the chemical potential difference and the electrical potential difference across a membrane, driven by the movement of protons.
Chemical Potential Difference: The Concentration Aspect
At its most fundamental level, the concentration gradient is the difference in the concentration of protons across a membrane. In cellular respiration, protons are actively pumped from the mitochondrial matrix to the intermembrane space, creating a higher concentration of H⁺ ions in the latter. Conversely, in photosynthesis, protons are pumped from the stroma into the thylakoid lumen, leading to a higher concentration within the lumen. This difference in proton concentration creates a tendency for protons to flow down their concentration gradient, from an area of high concentration to an area of low concentration. This inherent tendency to move from high to low concentration is a fundamental principle of thermodynamics and represents a form of potential energy.
Electrical Potential Difference: The Charge Aspect
In addition to the concentration difference, there is also an electrical potential difference across the membrane. Protons are positively charged ions. When they accumulate on one side of the membrane, they create an excess of positive charge on that side relative to the other. This charge separation generates an electrical potential, or voltage, across the membrane. Protons, being positively charged, are repelled by the excess positive charge on the side where they are concentrated and attracted to the relatively more negative charge on the opposite side. This electrical gradient further contributes to the driving force for proton movement.
The Synergy of Chemical and Electrical Gradients
The proton-motive force is the sum of these two forces. It represents the total potential energy available to drive cellular processes. While the chemical gradient is a significant factor, the electrical gradient can also play a substantial role, especially when the concentration difference is moderate. The PMF is typically expressed in millivolts (mV) and quantifies the overall electrochemical potential that can be harnessed by ATP synthase. This force is so potent that it can drive the synthesis of multiple ATP molecules per single pass of protons through the enzyme.
The Rotary Mechanism: Harnessing the Gradient
The elegance of ATP synthase lies in its ability to couple the flow of protons down their electrochemical gradient to the mechanical rotation of its subunits, which in turn drives the catalytic synthesis of ATP.
The F₀ Rotor: The Proton Channel

The F₀ sector of ATP synthase serves as a transmembrane channel through which protons can flow. This sector is composed of several subunits, including the ‘c’ ring and the ‘a’ subunit. The ‘c’ ring is embedded within the membrane and rotates as protons pass through. Each ‘c’ subunit typically has a proton-binding site. When a proton binds to a site on the ‘c’ subunit in the higher-concentration side of the membrane, it neutralizes the charge, allowing that subunit to rotate away from the inhibitory interaction with the ‘a’ subunit. As the ‘c’ ring rotates, it moves through the membrane.
On the other side of the membrane, in the lower-concentration region, the proton is released from the binding site. This release is facilitated by a specific amino acid residue in the ‘a’ subunit that provides a hydrophilic pathway for proton release. This cyclical binding and release of protons to and from the ‘c’ ring subunits is what drives the continuous rotation of the entire ‘c’ ring. The number of ‘c’ subunits in the ring can vary between species, influencing the number of protons required to complete one full rotation and, consequently, the number of ATP molecules synthesized per rotation.
The F₁ Motor: ATP Synthesis in Action
The F₁ sector is the catalytic headpiece of ATP synthase, responsible for the actual synthesis of ATP from adenosine diphosphate (ADP) and inorganic phosphate (Pi). It is attached to the F₀ sector via a central stalk, which is itself composed of the ‘γ’ and ‘ε’ subunits. This central stalk is rigidly connected to the rotating ‘c’ ring of the F₀ sector. Therefore, as the ‘c’ ring rotates, the central stalk also rotates, causing conformational changes in the catalytic sites located in the hexameric ring of α and β subunits within the F₁ sector.
Conformational Changes and Catalysis
The F₁ sector contains three catalytic sites, arranged in an alternating αβ configuration. Each site cycles through three distinct conformations: “Open” (O), “Loose” (L), and “Tight” (T).
- Open (O): In this state, the catalytic site has a low affinity for ADP and Pi and is ready to bind them.
- Loose (L): After ADP and Pi bind, the site transitions to the Loose conformation, where they are held loosely but are correctly positioned for reaction.
- Tight (T): The rotation of the central stalk induces the Tight conformation, which binds ADP and Pi very tightly. This tight binding strains the bonds within ADP and Pi, facilitating their condensation into ATP. The energy released from the proton gradient is used to drive this conformational change and the subsequent release of ATP.
The rotation of the central stalk, powered by the electrochemical gradient, forces each catalytic site through these three states sequentially. For every three protons that flow through the F₀ sector, one full rotation of the central stalk occurs, and each catalytic site synthesizes one molecule of ATP. This elegant coupling of ion flow to mechanical rotation and then to chemical synthesis is a testament to the efficiency of biological systems.
Beyond Protons: Sodium-Motive Force and Other Gradients
While the proton-motive force is the most well-studied and prevalent driving force for ATP synthesis, some organisms utilize other ions to generate the necessary electrochemical gradient.
Sodium-Motive Force (SMF)
Certain bacteria, particularly those living in environments with high salt concentrations or those involved in specific metabolic processes, employ a sodium-motive force (SMF) to power ATP synthesis. In these cases, sodium ions (Na⁺) are pumped across the membrane, creating a concentration and electrical gradient for Na⁺. ATP synthase in these bacteria, often referred to as Na⁺-ATP synthase, utilizes the flow of sodium ions instead of protons to drive its rotary mechanism and produce ATP. This adaptation allows these organisms to thrive in conditions where maintaining a proton gradient might be less efficient or energetically costly.

Other Ionic Gradients
While less common, research has suggested that other ionic gradients, such as those involving potassium ions (K⁺) or even calcium ions (Ca²⁺), might, under specific circumstances or in specialized organisms, contribute to or even drive ATP synthesis. However, these remain more specialized cases compared to the ubiquitous proton- and sodium-motive forces. The underlying principle remains the same: harnessing the potential energy stored in an electrochemical gradient of charged ions to power the molecular machinery of ATP synthase.
The discovery and elucidation of the mechanisms by which concentration gradients power ATP synthase represent a monumental achievement in biochemistry and biophysics. This fundamental process is not only essential for the survival of all known life forms but also serves as a powerful model for understanding how biological systems can efficiently convert potential energy into chemical energy, a principle that continues to inspire innovation in fields ranging from bioengineering to renewable energy technologies. The subtle yet mighty force of a concentration gradient, driving the relentless rotation of ATP synthase, underscores the profound elegance and efficiency of nature’s design.
